Web services
- GS-SMD: GS-SMD server for unfolding of peptide substrates in the active site of the γ-secretase complex
References:
U. Orzeł, P. Pasznik, P. Miszta, M. Lorkowski, S. Niewieczerzał, J. Jakowiecki, S. Filipek. GS-SMD server for steered molecular dynamics of peptide substrates in the active site of the γ-secretase complex. Nucleic Acids Res. (2023) 51, W251-W262. doi:10.1093/nar/gkad409
- GPCRsignal: web server for analysis of the interface between G protein-coupled receptors and their effector proteins by dynamics and mutations.
References:
P. Miszta, P. Pasznik, S. Niewieczerzał, J. Jakowiecki, S. Filipek. GPCRsignal: webserver for analysis of the interface between G-protein–coupled receptors and their effector proteins by dynamics and mutations, Nucleic Acids Res. (2021) doi:10.1093/nar/gkab434
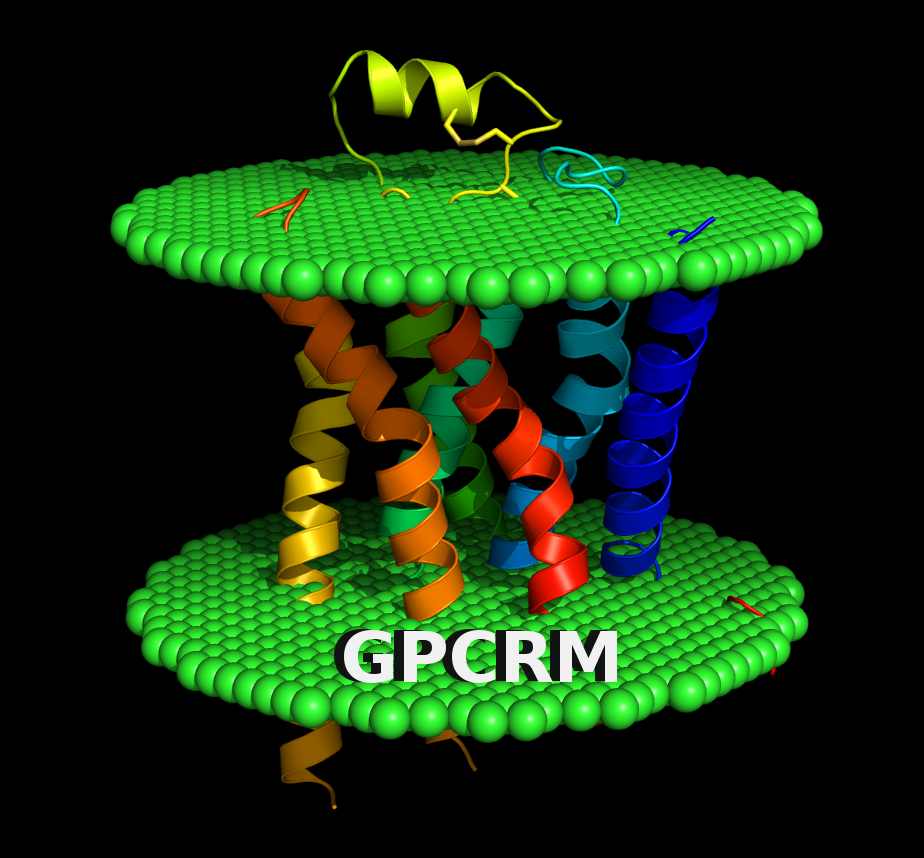
- GPCRM - The GPCR structure Modeling server.
References:
P. Miszta, P. Pasznik, J. Jakowiecki, A. Sztyler, D. Latek, S. Filipek, GPCRM: a homology modeling web service with triple membrane-fitted quality assessment of GPCR models, Nucleic Acids Research (2018), Web Server Issue (W1), doi:10.1093/nar/gky429
D. Latek, P. Pasznik, T. Carlomagno, S. Filipek (2013) Towards improved quality of GPCR models by usage of multiple templates and profile-profile comparison.
PLOS ONE 8(2): e56742, doi:10.1371/journal.pone.0056742
THE LATEST NEWS
- July 2023.GS-SMD web server
for SMD simulations of γ-secretase complex.
Publication in Nucleic Acids Research 2023 Web server issue.
- August 2022.COGRIMEN
- June 2021.GPCRsignalOur new service GPCRsignal was recently published in NAR 2021, W1.